Authors: Carlos Eduardo González-Penagos, Jesús Alejandro Zamora-Briseño, Monica Améndola-Pimenta, José Miguel Elizalde-Contreras, Flor Árcega-Cabrera, Yanis Cruz-Quintana, Ana María Santana-Piñeros, Mayra Alejandra Cañizárez-Martínez, Juan Antonio Pérez-Vega, Eliel Ruiz-May, Rossanna Rodríguez-Canul
https://doi.org/10.1016/j.taap.2022.116033
Abstract
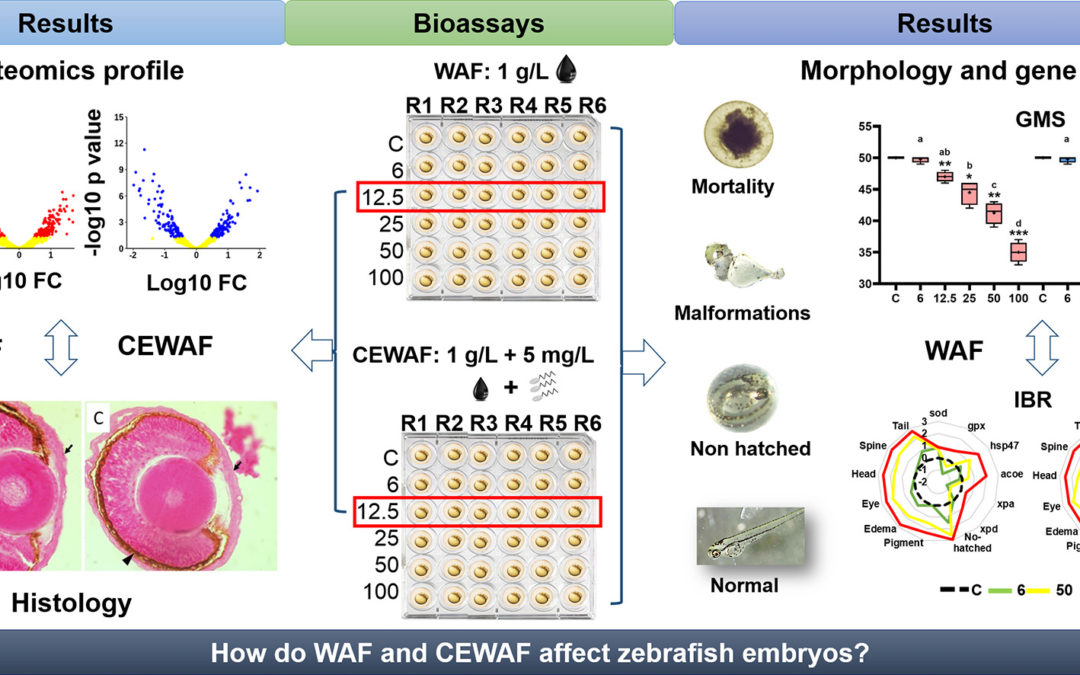
The effects of crude oil spills are an ongoing problem for wildlife and human health in both marine and freshwater aquatic environments. Bioassays of model organisms are a convenient way to assess the potential risks of the substances involved in oil spills. Zebrafish embryos (ZFE) are a useful to reach a fast and detailed description of the toxicity of the pollutants, including both the components of the crude oil itself and substances that are commonly used for crude oil spill mitigation (e.g. surfactants). Here, we evaluated the survival rate, as well as histological, morphological, and proteomic changes in ZFE exposed to Water Accumulated Fraction (WAF) of light crude oil and in mixture with dioctyl sulfosuccinate sodium (DOSS, e.g. CEWAF: Chemically Enhanced WAF), a surfactant that is frequently used in chemical dispersant formulations. Furthermore, we compared de hydrocarbon concentration of WAF and CEWAF of the sublethal dilution. In histological, morphological, and gene expression variables, the ZFE exposed to WAF showed less changes than those exposed to CEWAF. Proteomic changes were more dramatic in ZFE exposed to WAF, with important alterations in spliceosomal and ribosomal proteins, as well as proteins related to eye and retinal photoreceptor development and heart function. We also found that the concentration of high molecular weight hydrocarbons in water was slighly higher in presence of DOSS, but the low molecular weight hydrocarbons concentration was higher in WAF. These results provide an important starting point for identifying useful crude-oil exposure biomarkers in fish species.
Keywords: WAF/CEWAF, Malformations, Gene expression, Histology, Proteomics

Recent Comments