Autores: Rodríguez-Salazar Julieta, Almeida-Juarez Arisbeth G., Ornelas-Ocampo Katya, Millán-López Sofía, Raga-Carbajal Enrique, Rodríguez-Mejía José Luis, Muriel-Millán Luis Felipe, Godoy-Lozano E. Ernestina, Rivera-Gómez Nancy, Rudiño-Piñera Enrique, Pardo-López Liliana
https://doi.org/10.3389/fmicb.2020.01100
Abstract:
Catechol 1,2 dioxygenases (C12DOs) have been studied for its ability to cleavage the benzene ring of catechol, the main intermediate in the degradation of aromatic compounds derived from aerobic degradation of hydrocarbons. Here we report the genome sequence of the marine bacterium Pseudomonas stutzeri GOM2, isolated from the southwestern Gulf of Mexico, and the biochemical characterization of its C12DO (PsC12DO). The catA gene, encoding PsC12DO of 312 amino acid residues, was cloned and expressed in Escherichia coli. Many C12DOs have been described as dimeric enzymes including those present in Pseudomonas species. The purified PsC12DO enzyme was found as an active trimer, with a molecular mass of 107 kDa. Increasing NaCl concentration in the enzyme reaction gradually reduced activity; in high salt concentrations (0.7 M NaCl) quaternary structural analysis determined that the enzyme changes to a dimeric arrangement and causes a 51% decrease in specific activity on catechol substrate. In comparison with other C12DOs, our enzyme showed a broad range of action for PsC12DO in solutions with pH values ranging from neutral to alkaline (70%). The enzyme is still active after incubation at 50°C for 30 min and in low temperatures to long term storage after 6 weeks at 4°C (61%). EDTA or Ca2+ inhibitors cause no drastic changes on residual activity; nevertheless, the activity of the enzyme was affected by metal ions Fe3+, Zn2+ and was completely inhibited by Hg2+. Under optimal conditions the kcat and Km values were 16.13 s–1 and 13.2 μM, respectively. To our knowledge, this is the first report describing the characterization of a marine C12DOs from P. stutzeri isolated from the Gulf of Mexico that is active in a trimeric state. We consider that our enzyme has important features to be used in environments in presence of EDTA, metals and salinity conditions.
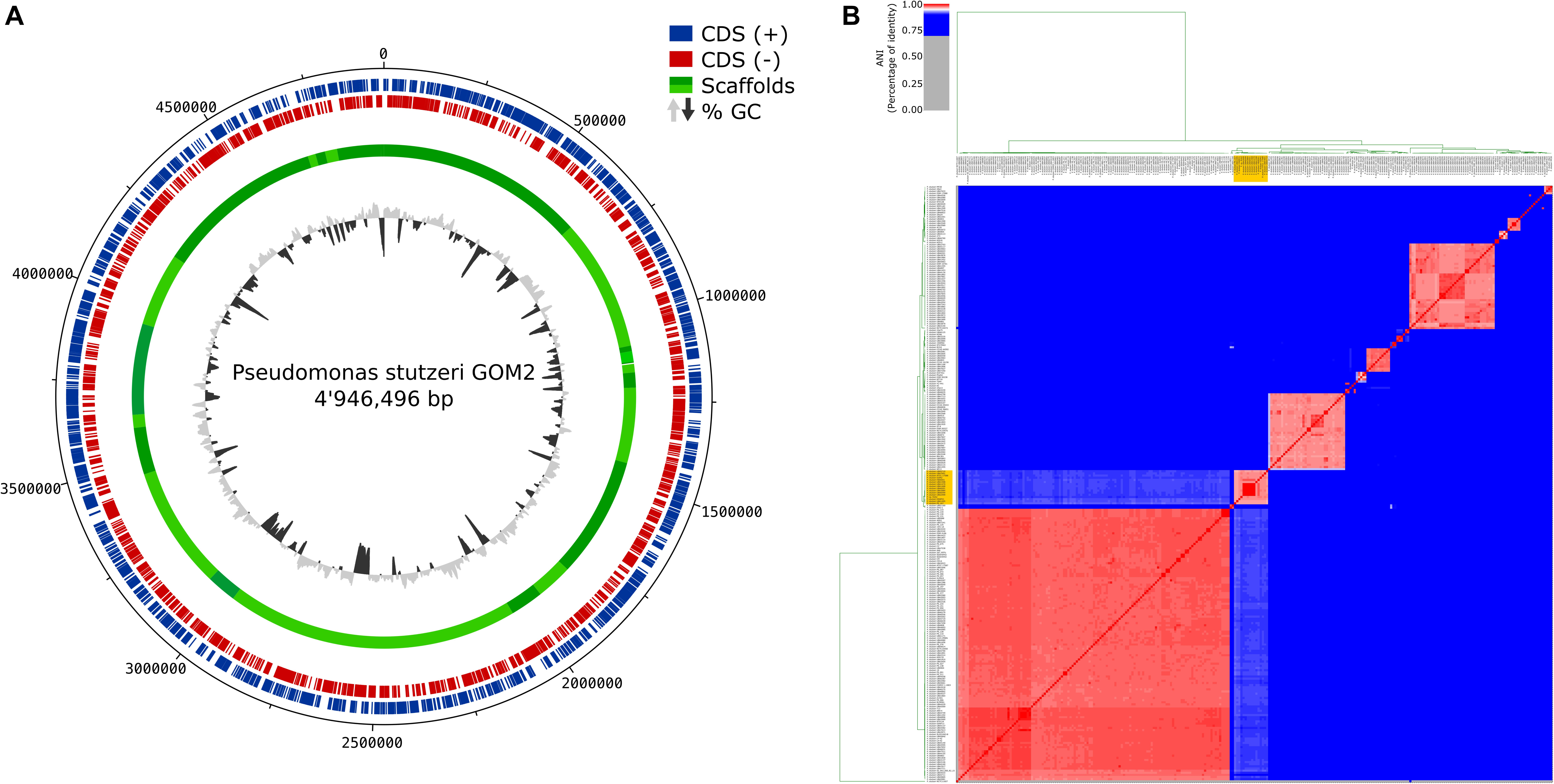
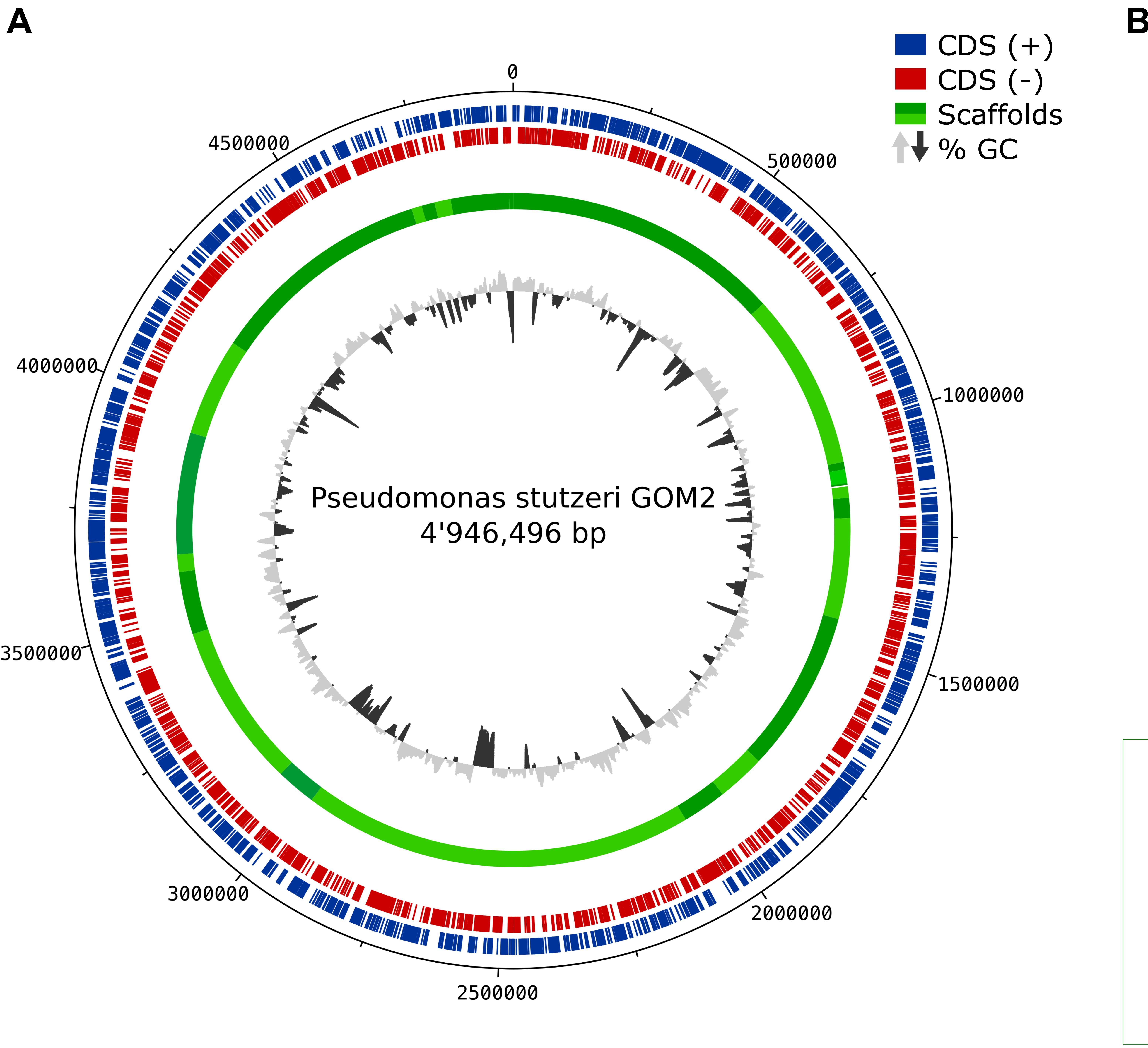
Figure 1. Genome assembly and ANI comparison. (A) Circular diagram of P. stutzeri GOM2 showing (from inner to outer)% G + C, GC skew and scaffolds. (B) Average nucleotide identity heatmap with all the available P. stutzeri genomes from GenBank. The yellow box indicates the group to which P. stutzeri GOM2 is related.

Recent Comments